Bio#
Research interests#
I work on molecular tool development for zebrafish neuroscience, the genetic identity of functionally identified neurons, and depth perception in larval zebrafish.
Molecular tool development#
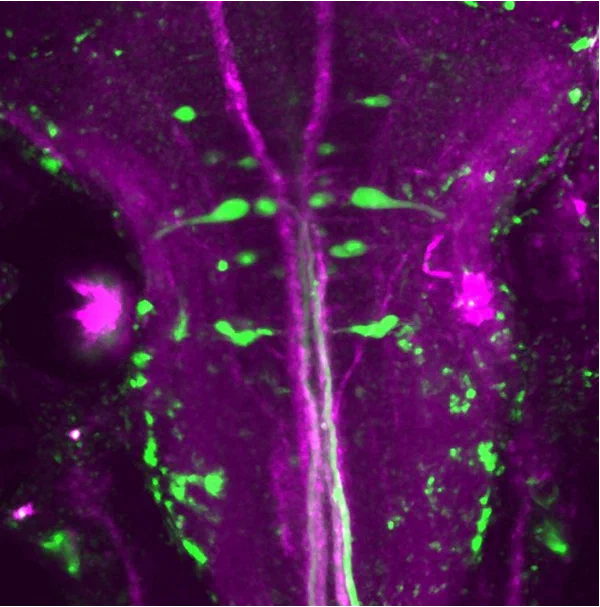
In our lab, we functionally identify neurons using whole-brain calcium imaging. However, subsequent experiments, such as morphological characterization or functional manipulation of individual neurons, require specific molecular toolkits. Existing driver lines in zebrafish lack the necessary specificity to dissect the circuits we study.
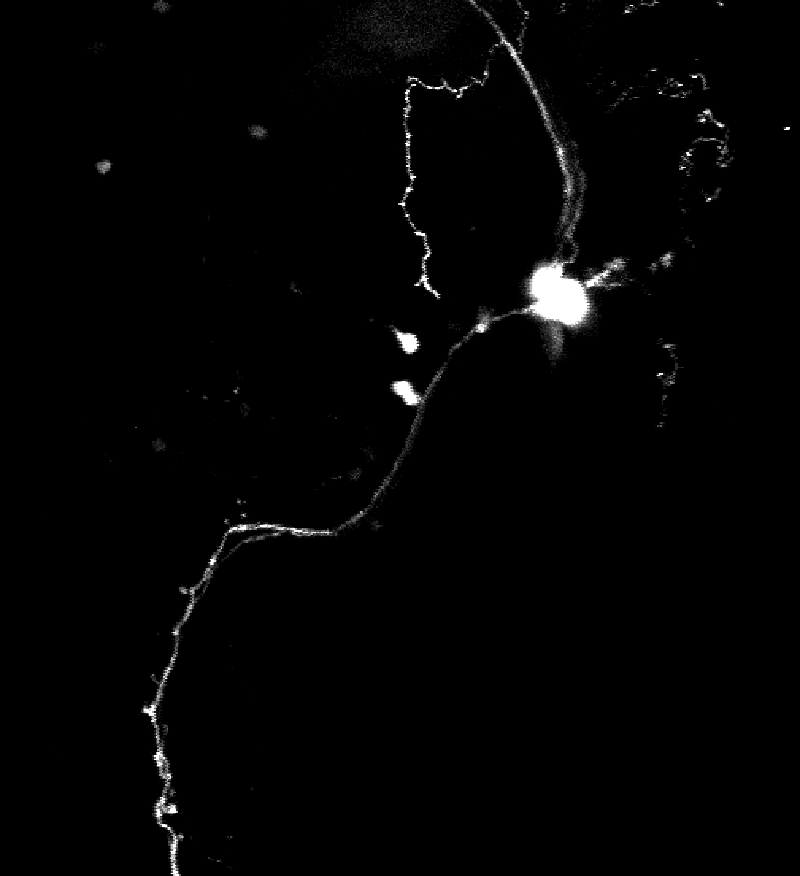
My work therefore focuses on developing light-inducible systems that enable single-cell precision targeting. These tools must be compatible with our calcium imaging pipeline; in particular, they should not rely on green fluorescence and must be activatable using a two-photon setup. To meet these requirements, I focus on UV-activatable systems combined with red fluorescent reporters.
Genetic identity of neurons#
To gain deeper insight into the molecular mechanisms of the neurons we study, we employ transcriptomic approaches. In particular, I use the molecular tools I develop to enable single-cell sequencing of functionally identified neurons.
In parallel, we collaborate with international partners on spatial transcriptomics projects to preserve anatomical context and link molecular profiles to neural circuits.
Motion parallax#
I am interested in how larval zebrafish perceive depth, particularly along the z-axis. My research focuses on motion parallax, that is, how larval zebrafish use self-generated movement to extract depth cues from their environment.
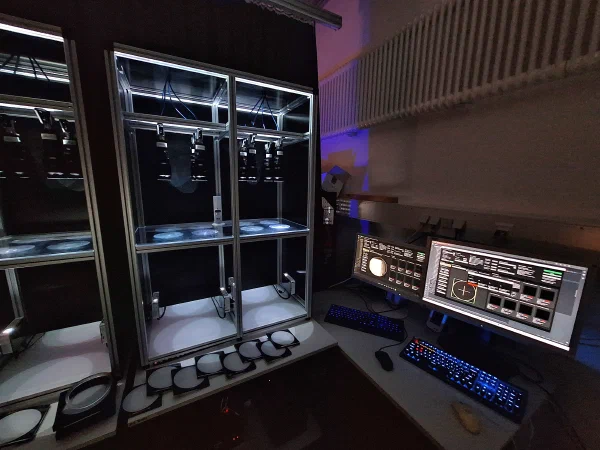
Using our virtual reality setup, I present the fish with different underlying geometries, analogous to a human walking across a glass floor and perceiving structures below. In parallel, I am developing virtual fish models that replicate the behavior of real fish under these conditions, with the goal of identifying neural correlates of the underlying computational components.
Additionally, I am developing a 3D tracking system for larval zebrafish, which will enable experiments in a full water tank rather than a constrained dish environment.
Miscellaneous#
I have a background in cell biology, with experience in molecular biology, genetics, and live-cell imaging in yeast. Transitioning to molecular and computational neuroscience in zebrafish was challenging, but ultimately very rewarding. I am motivated to continuously expand my skill set and to support my lab mates wherever I can.
I try to approach setbacks with perspective, maintain a sense of humor in difficult situations, and value engaging discussions, whether over coffee or a beer.
Projects#
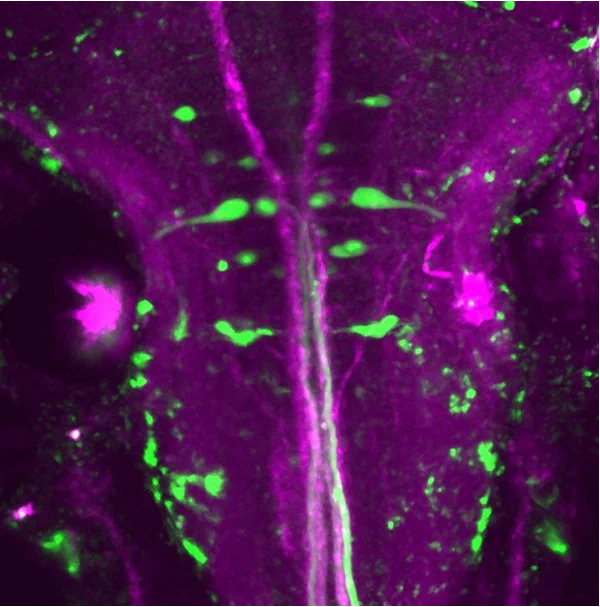
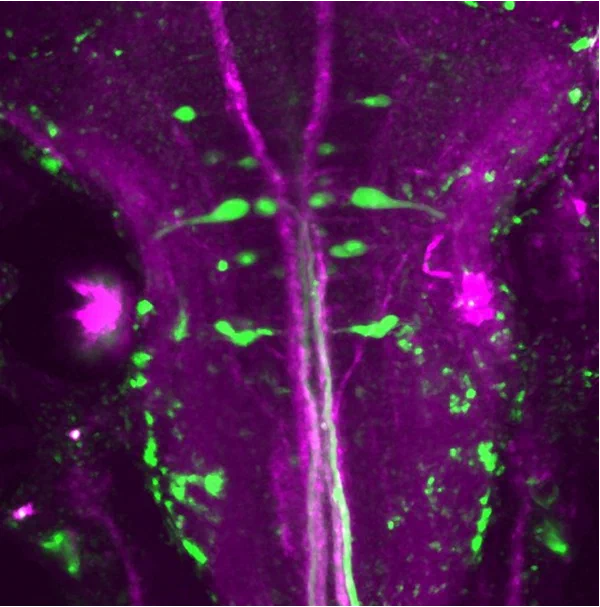
Molecular identity of neuronal structure and function
Methods#
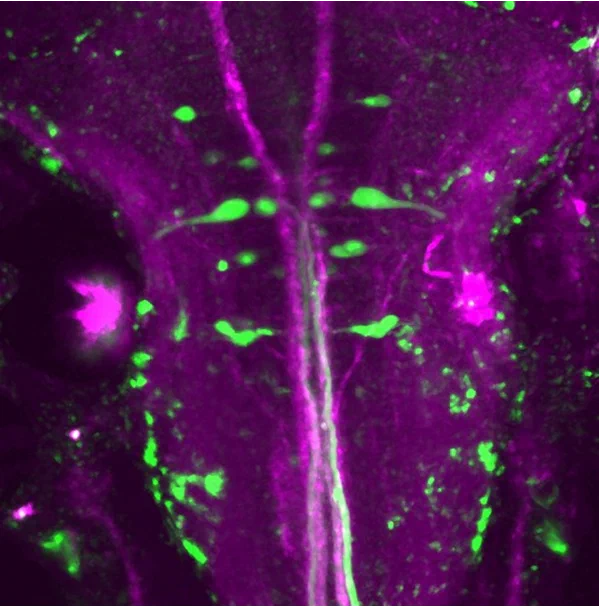

Thesis#
CV#
Positions
Education
Publications#
2023
- Hummel D. R., Hakala M., Toret C. P., Kaksonen M. (2023) Bsp1, a fungal CPI motif protein, regulates actin filament capping in endocytosis and cytokinesis. https://doi.org/10.1101/2023.10.09.561521
- Hummel D. R., Kaksonen M. (2023) Spatio-temporal regulation of endocytic protein assembly by SH3 domains in yeast. Molecular Biology of the Cell. https://doi.org/10.1091/mbc.e22-09-0406
2022
- Hummel D. (2022) Regulation of membrane scission in yeast endocytosis. Molecular Biology of the Cell. https://doi.org/10.1091/mbc.e21-07-0346
- Hummel D. R., Kaksonen M. (2022) Spatio-temporal regulation of endocytic protein assembly by SH3 domains in yeast. https://doi.org/10.1101/2022.08.11.503583
2021
- (2021) The Gb3-enriched CD59/flotillin plasma membrane domain regulates host cell invasion by Pseudomonas aeruginosa.. Cellular and molecular life sciences : CMLS. https://doi.org/10.1007/s00018-021-03766-1
2018
- (2018) In Vitro and in Cellulo Sensing of Transition Metals Using Time-Resolved Fluorescence Spectroscopy and Microscopy.. Journal of fluorescence. https://doi.org/10.1007/s10895-018-2335-z
